Overview
- Products: Online open-source portal for genotypes and phenotypes.
- Cost: Free
- Reports: Report on genotype and phenotype
- Raw data access: OpenSNP does not offer DNA testing and requires users to have access to their raw DNA data.
- Privacy: Once the user uploads their data, it is open for all to access.
- Alternatives: SelfDecode – The best option for health-focused DNA analysis with personalized reports, symptom analysis, and health recommendations.
Pros
- Open library for both public and health professionals on genotype and phenotype
- Free to use
- Easy to understand reports and downloadable datasets
Cons
- Once you upload your information, it is no longer private
- The datasets may not yet be 100% accurate
- Does not provide any health recommendations based on DNA
About OpenSNP
OpenSNP is a website where users upload their genetic information. This open-source site allows users to share their personal information, such as medical history and genetic information.
Founded by Bastian Greshake Tzovaras in 2011, this community-driven and community-owned project is maintained by him along with Helge Rausch and Philipp Bayer.
The project isn’t associated with any institution and is fueled by the contribution of its members. It is a patient-led research (PLR) program that could redefine the way we conduct health research.
Review of OpenSNP Products & Features
OpenSNP allows users to share their test results with others while simultaneously allowing them to find other people with similar genetic variations. This makes for a great online library of primary literature on genetic variations that scientists can use.
Genotyping Users
The following features are available for genotyping users.
Upload Data
People who were genotyped from 23andMe, deCODEme, or FamilyTreeDNA can share their results with the world. All they have to do is upload the raw genotyping or exome data on the website. They can also use it as a personal profile for their genetic test results.
Share Phenotypes
By sharing phenotypes with other users, people that have similar characteristics and traits can connect with one another. Not only can this help with networking, but most importantly, this will help scientists to discover new genetic variations and information.
Share Stories
Users can share stories about their genetic variations and phenotypes on the OpenSNP website. Likewise, they can read what other users have to say, and this ultimately creates a community of shared knowledge.
Find Literature On Genetic Variation
Genotyping users can find the latest open-access journal articles on genetic variations. Popular articles from the Public Library of Science are available. In case anyone finds it difficult to understand, SNPedia summarizes the articles for easier comprehension.
Scientists
OpenSNP has features for scientists as well.
Search for Phenotypes
Many diseases and traits are assumed to possess genetic components. Using genetic markers, scientists can find samples for research. If a genetic marker isn’t present, they can add it to the website for development consideration.
Easily Download Datasets
Scientists can easily download mass collections of genotyping raw data. The files can also be grouped according to the genetic variations based on specific phenotypes. Datasets from 23andMe, deCODEme, and FamilyTreeDNA can be easily collected for association studies.
Get Notified About New Data
Scientists don’t need to check for new entries. There is a feed for new datasets and therefore, scientists can easily get all the new data for the phenotype of their interest.
Review of OpenSNP Reports
The website allows anyone to participate in any stage of genomics research. The proof-of-concept mechanism makes it easy for any user to join.
Given that the project isn’t associated with any government organization or any other institution or organization for that matter, anyone has the opportunity to be a part of the extensive research program.
This act of sharing data adds value to the users’ contribution and shows how a community can get together for the good of research. Moreover, the transparent open-source code makes the data accessible to anyone. This invites greater scrutiny and oversight than other closed-source programs.
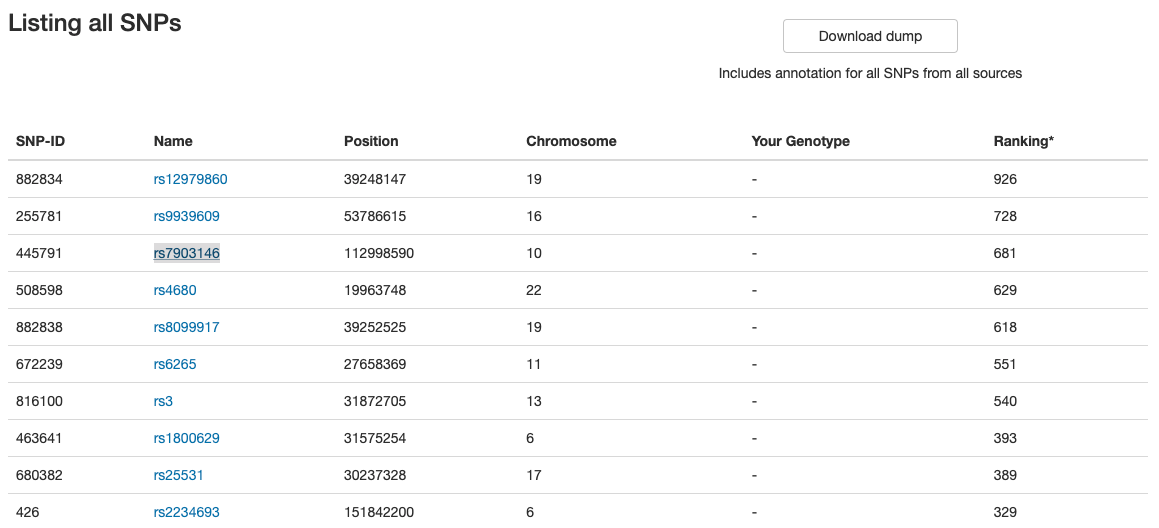
OpenSNP offers a list of SNPs open for anyone to access. The website does not require an account to view or download any of the information available. The screenshot below shows a list of SNPs:
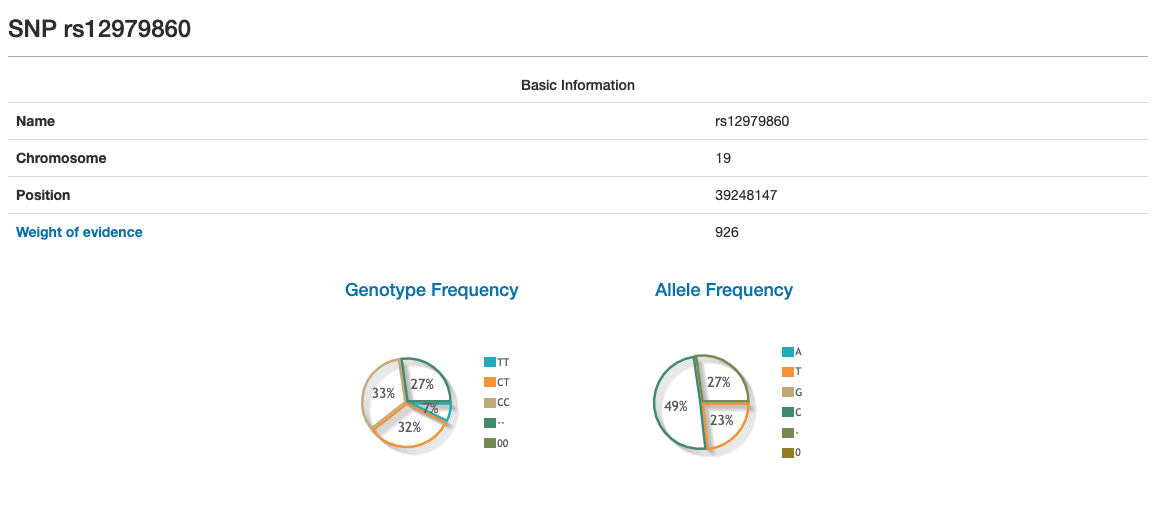
By clicking on any link from the Name column, OpenSNP sends the user to another page with more information about each SNP.
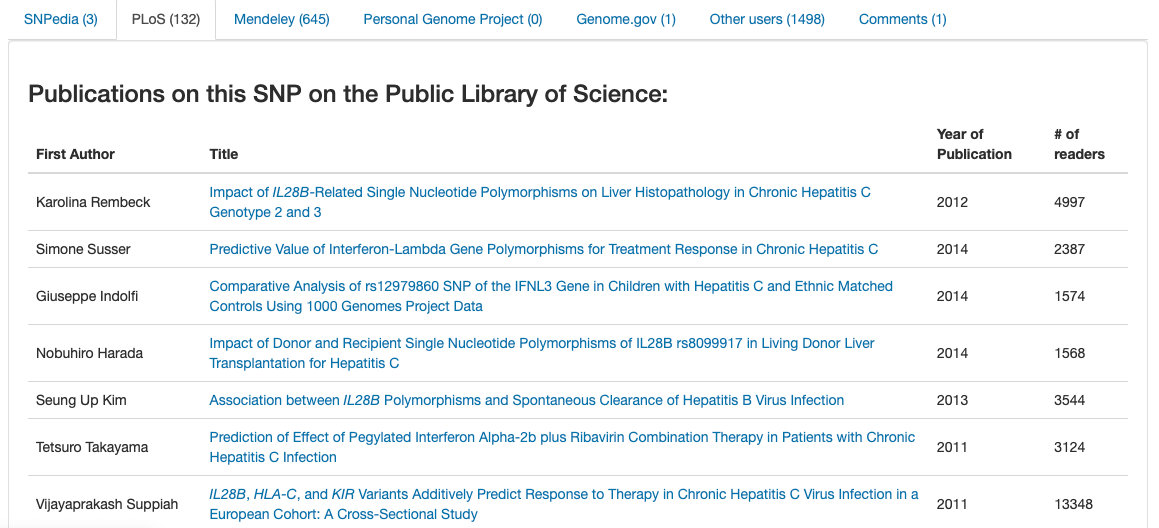
Furthermore, the website provides an easy way to research and access scientific publications related to the variant. As seen on the screenshot below, OpenSNP informs users of the details of each publication such as the author’s name, year of publication, and the number of readers.
SNPedia also summarizes the information of articles making it easier for users to comprehend.
Furthermore, the categorization of data according to phenotypes makes them convenient for anyone to study.
Cost of OpenSNP
The founders of OpenSNP took the “open” in their name quite literally. When they say the data is open, they mean the datasets are accessible to anyone and everyone for free.
Bastian was fed up when he found a lack of genotyping data on the internet. As a result, he launched OpenSNP to encourage people to share their genetic data to further research studies. The goal wasn’t to make money but to make science more open and accessible.
Users can simply sign up at the website and choose to upload their data, or even download the datasets arranged in the site’s data section.
Health Recommendations from OpenSNP
OpenSNP doesn’t provide feedback to users based on their test reports. However, given that this is a community-driven project, there is potential for improvements. Given that OpenSNP is a patient-led research program, we may see new initiatives as the project matures.
Review of OpenSNP Privacy & Data Security
Unlike many websites, OpenSNP openly declares that the information a user uploads on their website is not private, and may be forwarded to third parties if need be, to help research.
Even if a user deletes their information, they will not be able to completely get rid of the data, as any of the information backed up in the system may have already been downloaded and published elsewhere.
OpenSNP asks users twice for their confirmation, once while signing up on the website, and again when uploading data.
The website is licensed under Creative Commons Zero, meaning no rights are reserved on the data once it’s uploaded. After someone’s personal information or genetic report has been published, it is accessible to the world.
They make it clear that the goal of the project is not to sell user data but to make science more open and accessible.
SelfDecode vs OpenSNP
- SelfDecode delivers natural supplement, diet, and lifestyle suggestions based on your genes that you can implement right away. OpenSNP does not offer any health suggestions based on your genetic data.
- SelfDecode tells you why they make each recommendation so that you can understand the science behind the suggestion. OpenSNP only gives you information about users’ genetic information.
- SelfDecode prioritizes recommendations based on their analysis of all the relevant genes instead of one gene at a time (through reports). OpenSNP does not offer any health recommendations.
- SelfDecode takes a holistic approach to give recommendations that are best for your genes AND the health topic. OpenSNP does not give recommendations to optimize your health.
- SelfDecode is the most comprehensive and looks at more genes & SNPs (over 700,000) to deliver the best analysis of genetic risks. OpenSNP analyzes your whole DNA file, which is limited to the SNPs included in a particular company’s test. They do not offer DNA testing.
- SelfDecode supports everything with peer-reviewed scientific studies in their research and checks for contradicting information. OpenSNP references scientific articles written about certain genes and the alleles involved in determining traits.
- SelfDecode never sells your data or gives it away. OpenSNP is a crowdsourced and completely open database.
Comparisons
|
SelfDecode |
OpenSNP | Gene Heritage |
Promethease |
|
| Personalized & holistic health recommendations |
Yes |
No | No |
No |
| Cost (USD) |
$97 – $389 |
Free | $12 per family member |
$12 |
| Products |
DNA testing, wellness reports, research-based personalized blog posts, health recommendations |
Online open-source portal for genotypes and phenotypes. | DNA analysis, Individual, Parent-Child, and Grandchild trait reports |
DNA analysis, health reports |
| Raw data access |
Yes |
N/A | N/A |
N/A |
OpenSNP Reviews
There are no customer or user reviews online, but the OpenSNP Wikipedia entry lays out the potential risks and benefits. They also have entries in online journals such as NCBI, ResearchGate, and Plos One which contain detailed information about the platform and its features.
Alternatives to OpenSNP
SelfDecode: The best option for health-focused DNA analysis with personalized reports and recommendations to improve your quality of life.
OpenSNP Review Summary
OpenSNP allows users to share their test results for free with others while also allowing them to find other people with similar genetic variations. This makes for a great online library of primary literature on genetic variations that scientists can use.
But this openness may come at a cost, and there are still many bioethical considerations to think through. The biggest point of debate is informed consent. It is arguable whether participants of projects such as openSNP have understood the potential risks they take by making their data available. While users can participate with a pseudonym, it will always be possible to identify them because of the unique nature of an individual’s genome.
For those seeking DNA-based insights to help them improve their health, OpenSNP might not be the best option. The company does offer access to published scientific research on genetic variations, but finding relevant articles and understanding them might require some previous knowledge.
An alternative such as SelfDecode offers a DNA test to deliver easy-to-understand diet, lifestyle, and supplement recommendations aimed at improving overall well-being. SelfDecode takes privacy very seriously and will not sell or share your data with anyone. For those scientifically curious, SelfDecode embeds all scientific references used throughout the reports.
Related
- Gene Heritage Review: A Simple Insight Into Your Genetics
- Promethease Review: What can DNA research tell us?